Regulation of mRNA translation
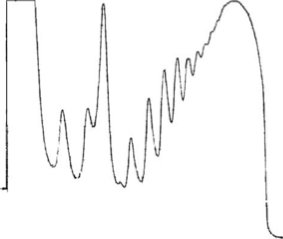
Canonical initiation involves the recruitment of the 40S ribosome to the 5' end of the mRNA. The 40S then scans through the 5'-UTR to find the start codon, where the 60S joins and elongation begins. We are interested in how translation is regulated during initiation, especially by the 5'-UTR and associated proteins. Disruptions to RNA-protein interactions and translational regulation play significant roles in a variety of cancers and other disorders (e.g. spinal muscular atrophy).
|
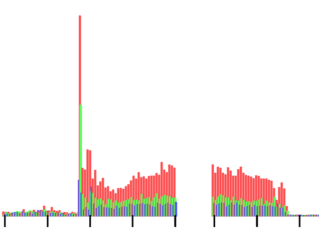
Function of cytoplasmic long non-coding RNAsWhilst the majority of our genome is transcribed, only a small fraction is protein-coding. This implies that many non-coding RNAs exist, some of which are similarly processed to mRNAs, termed long non-coding RNAs (lncRNAs). The majority of characterised lncRNAs are nuclear but recent ribosome profiling results have revealed that many are translated. Although controversial, it is clear that numerous lncRNAs are cytoplasmic, polysome-associated and translated. The line between coding and non-coding has become blurred and some lncRNAs are really mRNAs. LncRNAs are enriched in neuronal tissues and several have been found associated with neurological conditions e.g. Alzheimer’s disease. To better understand the molecular basis of neuronal disease it is vital that we appreciate the role of cytoplasmic lncRNAs in regulating gene expression. We are studying the function of these cytoplasmic lncRNAs.
|
Evolution of functional lncRNA-protein complexes
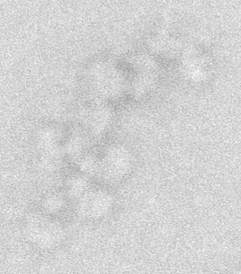
Although most lncRNAs are poorly conserved at the sequence level we can understand the importance and potential evolution of lncRNAs by studying them across multiple related organisms. We are interested in specific lncRNA-protein complexes and their conservation e.g. Xist. Our work seeks to understand the evolutionary relationship of lncRNAs and the proteins they interact with that are are important across numerous species. To dissect the molecular nature and function of these lncRNA-protein complexes we employ a combination of comparative genomics and biochemistry. We are also unravelling the relationship between lncRNA evolution, their protein-coding potential and molecular function.
Specialised ribosomes
|
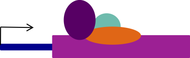
Until recently the dogma has been that ribosomes are homogenous macromolecular machines but it is now becoming clear that ribosomes exert specificity and regulatory capacity through variations in their protein and RNA composition. This allows the generation of different types of ribosomes, termed specialised ribosomes. One type of these specialised ribosomes contain paralogues of canonical ribosomal proteins, which exhibit tissue specific expression (e.g. RpL22-like in Drosophila testis). The function of these different ribosome versions are beginning to be elucidated with mutants of specific ribosomal proteins exhibiting precise phenotypes e.g. RPL38 mouse mutants exhibit reduced HOX gene translation and impaired neural specification. We aim to understand the structural and functional relationship of specialised ribosomes in Drosophila testes.
|